Connecting to IBM DB2 (JDBC)
This example illustrates how to connect to an IBM DB2 database server via JDBC.
Prerequisites
•JRE (Java Runtime Environment) or Java Development Kit (JDK) must be installed. This may be either Oracle JDK or an open source build such as Oracle OpenJDK. XMLSpy will determine the path to the Java Virtual Machine (JVM) from the following locations, in this order: (i) the custom JVM path you may have set in application Options; ; (ii) the JVM path found in the Windows registry; (iii) the JAVA_HOME environment variable.
•Make sure that the platform of XMLSpy (32-bit, 64-bit) matches that of the JRE/JDK.
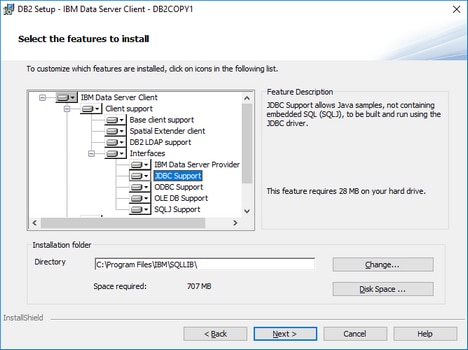
•The JDBC driver (one or several .jar files that provide connectivity to the database) must be available on your operating system. This example uses the JDBC driver available after installing the IBM Data Server Client version 10.1 (64-bit). For the JDBC drivers to be installed, choose a Typical installation, or select this option explicitly on the installation wizard.
If you did not change the default installation path, the required .jar files will be in the C:\Program Files\IBM\SQLLIB\java directory after installation.
•You have the following database connection details: host, port, database path, name, or alias, username, and password.
Connection
1.Start the database connection wizard and click JDBC Connections (see JDBC Connection for a screenshot of the dialog).
2.In the Classpaths field, enter the path to the .jar file that provides connectivity to the database. If necessary, you can also enter a semicolon-separated list of .jar file paths. If you have added the filepath to the CLASSPATH of the system, you can leave this field empty.
3.In the Driver field, select the appropriate driver. Relevant drivers will be available only if a valid .jar file path is in the Classpaths field or in the CLASSPATH environment variable.
4.Enter the username and password for the database in the corresponding fields.
5.Enter the JDBC connection string in the Database URL field according to the pattern in the table below, replacing the highlighted values with the ones applicable to your database server.
6.Click Connect.
Connection details of the IBM DB2 example
Field | Value |
Classpaths | C:\Program Files\IBM\SQLLIB\java\db2jcc.jar |
Driver | com.ibm.db2.jcc.DB2Driver |
Database URL | jdbc:db2://hostName:port/databaseName |